Migration - Analysis and modelling of microscopy images (large data sets)
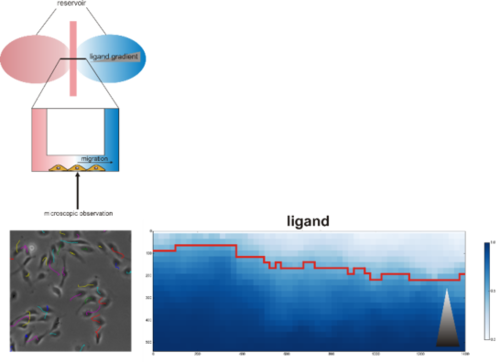
Top: Model of chemotaxis assay chamber (adapted from Ibidi). Left: Annotated trajectories for migrating cells. Right: Markov committor analysis of migratory behavior (gray triangle: growth factor gradient)
Directed cell migration is a complex process that involves cell polarization induced by chemical signals including growth factors, cell adhesion, formation of protrusions, retraction and contraction of a single cell or cell collective. The polarization depends on different mechanochemical signals including chemical compositions of the substrate, substrate stiffness, geometries and adhesive cues.
Migration is a highly coordinated process that involves synchronized rearrangements of cytoskeleton, dissolution of cell-cell contacts and polarization to orient the cell towards a targeted area.
The BMP signaling pathway is acknowledged to control migration of diverse cell types, including neural crest cells, mesenchymal progenitor cells and endothelial cells. The exact molecular mechanism is still unresolved. Cancer, hypoganglionosis, Hirschsprung's disease and intestinal neuronal dysplasia are pathologies with aberrant BMP signaling resulting in altered cell migration emphasizing the role of the pathway in migration.
Our lab investigates the mechanism by which BMPs induce (directional) migration/chemotaxis in various cell types and its functional implication in diseases. We use chemotaxis assays in which chemoattractant gradients are formed with simultaneous digital time lapse recording of cell movement to be able to track migration of cells in response to stimuli. Automated analysis of large data collections from these experiments are done in collaboration with the “Image analysis in biology and material science” at the Zuse Institute Berlin (ZIB) and enables us to track migration speed and orientation of single cells in response to chemical stimuli.