Disease-associated mutations in TGF-β receptors alter the functionality of associated regulators
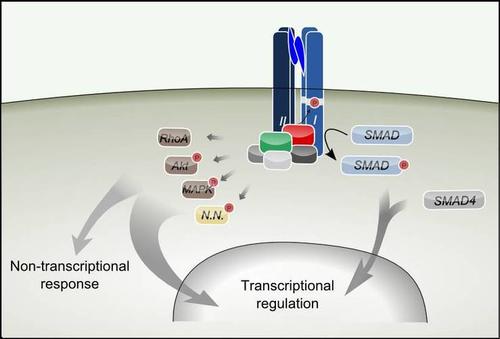
Fig. 1.TGF-beta superfamily ligands activate Smad and non-Smad pathways such as RhoA, Akt, and MAP Kinase (MAPK). There may be pathways which are yet to be discovered (N.N.).
Image Credit: Dr. Daniel Horbelt
Transforming growth factor-β (TGF-β) is a modulator in developmental processes and a regulator in homeostasis. Dysregulation of TGF-β signaling is associated with human diseases such as cancer and cardiovascular disease. Hereditary defects predispose to metabolic, musculoskeletal, and cardiovascular malfunctions.
Even though the principal functions and pathways of TGF-β signaling have been elucidated in the past decades, increasing levels of complexity regarding its role in vivo and its signaling mechanisms have emerged, emphasizing the highly context specific nature of this pathway.
TGF-β superfamily signaling is initiated by binding of an extracellular ligand to cognate receptors, which promotes the formation of active receptor signaling complexes. The signaling pathways that originate from these complexes, the canonical Smad pathway and several non-Smad pathways, trigger transcriptional regulation but also direct cellular responses (Fig. 1).
We are investigating the "signalotypes" of pathway components that exhibit an abnormal functionality as a consequence of disease-related inherited or somatic mutations. Mutations in TβRII (TGFBR2 gene) were found associated with a group of diseases that feature significant phenotypic similarity with Marfan syndrome (MFS). Our analysis of mutations in TβRII which are associated with Loeys-Dietz syndrome (LDS), type 2 Marfan syndrome (MFS2), or familial thoracic aortic aneurysms and dissections (FTAAD) revealed a correlation between clinical severity and signaling activity. Results from this study suggest that the balance of canonical TGF-ß-Smad signaling and non-canonical ERK signaling is a determinant for the development of thoracic aortic aneurysms and dissections (TAAD) on one hand and additional manifestations including skeletal and craniofacial phenotypes on the other hand.
The pathological consequences of mutant receptors, however, could not be fully explained to date on the basis of apparent defects in the receptor proteins themselves, such as altered expression, localisation, or enzymatic activity.
Thus, our most recent approaches to understand the consequences of pathological mutations in TGF-β receptors target levels further downstream, by identifying the differential interactomes of wildtype and mutant receptors. A series of receptor-interacting proteins have been identified which associate with the receptors in a context-dependent manner. Some of these proteins were demonstrated to modulate TGF-β signalling by either enhancing or repressing the signalling activity originating at the receptors. We have already generated the interactomes of various wildtype receptors of the TGF-β superfamily using different experimental strategies and have sucessfully identified and characterized novel regulators of TGF-β superfamily signalling.
By identifying proteins that selectively associate with wildtype or mutant TGF-β superfamily receptors, respectively, we will be able to elucidate mechanisms of disease development that are obscure to date.