Site-specific incorporation of fluorinated amino acids at protein-protein interfaces
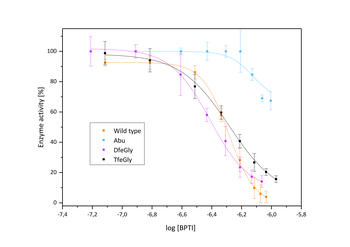
Inhibition of β-trypsin with BPTI variants
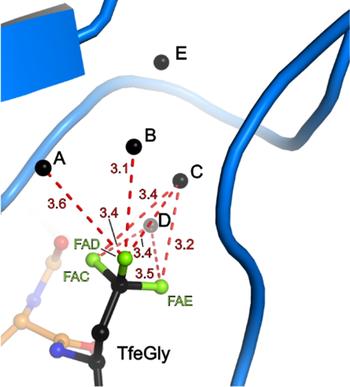
Binding pocket of β-trypsin-BPTI-TfeGly complex
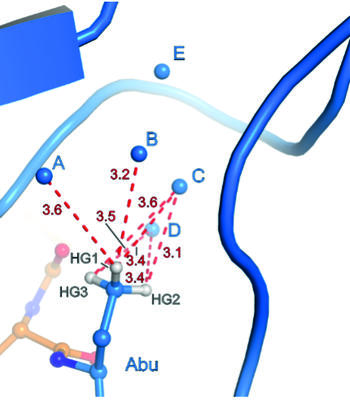
Binding pocket of β-trypsin-BPTI-Abu complex
Fluorinated amino acids have been incorporated into peptides in previous studies to investigate to what extent this unique element affects important properties, like biological activity and bioavailability. It has been shown that fluorination of the side-chains of otherwise canonical residues is in some cases advantageous, and typically does not perturb the native structure. Fewer studies have been undertaken with site-specific incorporation of fluorinated non-canonical amino acids derivatives into peptides. In general, the precise impact of fluorination is often difficult to predict and therefore requires further systematic investigation.
The presence of a single fluoroalkyl amino acid side chain (Figure, right panel) in the P1 position of the well-characterized serine-protease inhibitor BPTI (bovine pancreatic trypsin inhibitor) can fully restore inhibition properties (Figure, left panel) to a mutant containing the corresponding hydrocarbon side chain at the same position (Figure, center panel).
Further investigations revealing the impact of fluorine on protein-protein interactions are currently ongoing.
Further Reading
Ye, S.; Loll, B.; Berger, A. A.; Mülow, U.; Alings, C.; Wahl, M. C.; Koksch, B. Chem. Sci., 2015, 6, 5246-5254. (DOI:10.1039/C4SC03227F)