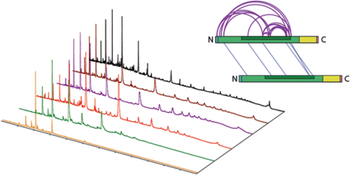
Oligomerisation of Synaptobrevin-2 Studied by Native Mass Spectrometry and Chemical Cross-Linking
Wittig, S.; Haupt, S.; Hoffmann, W.; Kostmann, S.; Pagel, K.; Schmidt, C.* – 2019
Synaptobrevin-2 is a key player in signal transmission in neurons. It forms, together with SNAP25 and Syntaxin-1A, the neuronal soluble N-ethylmaleimide-sensitive factor attachment protein receptor (SNARE) complex and mediates exocytosis of synaptic vesicles with the pre-synaptic membrane. While Synaptobrevin-2 is part of a four-helix bundle in this SNARE complex, it is natively unstructured in the absence of lipids or other SNARE proteins. Partially folded segments, presumably SNARE complex formation intermediates, as well as formation of Synaptobrevin-2 dimers and oligomers, were identified in previous studies. Here, we employ three Synaptobrevin-2 variants—the full-length protein Syb(1-116), the soluble, cytosolic variant Syb(1-96) as well as a shorter version Syb(49-96) containing structured segments but omitting a trigger site for SNARE complex formation—to study oligomerisation in the absence of interaction partners or when incorporated into the lipid bilayer of liposomes. Combining native mass spectrometry with chemical cross-linking, we find that the truncated versions show increased oligomerisation. Our findings from both techniques agree well and confirm the presence of oligomers in solution while membrane-bound Synaptobrevin-2 is mostly monomeric. Using ion mobility mass spectrometry, we could further show that lower charge states of Syb(49-96) oligomers, which most likely represent solution structures, follow an isotropic growth curve suggesting that they are intrinsically disordered. From a technical point of view, we show that the combination of native ion mobility mass spectrometry with chemical cross-linking is well-suited for the analysis of protein homo-oligomers.