Pre-mRNA splicing
Pre-mRNA splicing
Many eukaryotic precursor messenger RNAs (pre-mRNAs) bear coding regions (exons) interspersed with non-coding intervening sequences (introns). Production of mature mRNAs thus requires pre-mRNA splicing, during which introns are excised and neighboring exons are ligated. A splicing reaction consists of two SN2-type transesterification reactions. In the first half-reaction, the phosphodiester bond at the 5’-splice site is attacked by the 2’-hydroxyl of a conserved adenosine of the branch point sequence in the intron, generating a free 5’-exon and an intron lariat-3’-exon. In the second half-reaction, the 3’-hydroxyl of the 5’-exon attacks the phosphodiester bond at the 3’-splice site, leading to exon ligation and excision of the lariat intron. Splicing is carried out by spliceosomes, which are large and highly dynamic molecular RNP machines. Spliceosomes are composed by five small nuclear (sn) RNPs (U1, U2, U4, U5 and U6 in the case of the major/U2-type spliceosome) and many additional non-snRNP splicing factors. For each round of splicing, a spliceosome is assembled, catalytically activated and, after splicing catalysis, disassembled. Each assembly, activation, catalysis and disassembly step is associated with profound compositional and conformational remodeling of the underlying RNA-protein interaction network. Higher eukaryotic genes typically contain more than one intron and their pre-mRNAs can undergo alternative splicing, giving rise to mature transcripts with different combinations of exons or portions of exons and thus ultimately to more than one protein. Alternative splicing is essential for producing the large repertoires of proteins required in higher organisms from a limited number of protein-coding genes, endows eukaryotes with entirely novel mechanisms to regulate their gene expression and supports novel principles of evolution. Splicing and alternative splicing are also of tremendous medical interest, as mutations in splicing factor genes have been associated with more than 300 disorders in humans and as at least 25 % of all inherited human diseases root in aberrant splicing. We are interested in how the molecular design principles of spliceosomes ensure the reliable identification of authentic splice sites, yet at the same time provide sufficient functional flexibility for alternative splicing.
Recent publications
De Bortoli F, Neumann A, Kotte A, Timmermann B, Schüler T, Wahl MC, Loll B, Heyd F (2019) Increased versatility despite reduced molecular complexity – evolution, structure and function of metazoan splicing factor PRPF39. Nucleic Acids Res, doi: 10.1093/nar/gkz243. (PubMed)
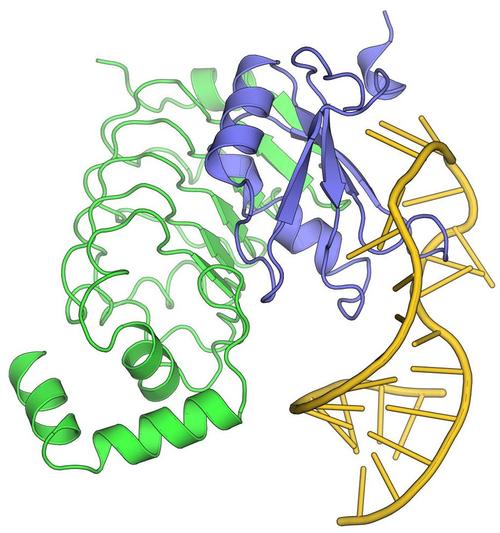
Weber G, DeKoster GT, Holton N, Hall K, Wahl MC (2018) Molecular principles underlying dual RNA specificity in the Drosophila SNF protein. Nat Commun 9, 2220. (PubMed)
Ulrich A, Seeger M, Schütze T, Bartlick N, Wahl MC (2016) Scaffolding in the spliceosome via single α-helices. Structure 24, 1972-1983. (PubMed)
Ulrich AKC, Schulz JF, Kamprad A, Schütze T, Wahl MC (2016) Structural basis for the functional coupling of the alternative splicing factors Smu1 and RED. Structure 24, 762-773. (PubMed)
Liu S, Mozaffari-Jovin S, Wollenhaupt J, Santos KF, Theuser M, Dunin-Horkawicz S, Fabrizio P, Bujnicki JM, Lührmann R, Wahl MC (2015) A composite, double-stranded/single-stranded RNA-binding region in the Prp3 protein important for U4/U6•U5 tri-snRNP stability and splicing. eLife 4, e07320. (PubMed)