Research
1. Developing superresolution microscopy as a readout of protein function
Most cellular functions are executed by the concerted interaction in time and space of several proteins in complexes. By combining several proteins with different functional domains, the gene products of a limited genome can execute the wealth of functions that is required to build and maintain higher functions of cell and organism.
While by now many individual protein-protein interactions are well understood, it is still unclear how multiprotein complexes are assembled from their components in cells and how their spatial organization relates to their function.
A novel approach to this problem may come from the recently developed single molecule localization-based superresolution-microscopy methods. These allow the determination of the exact position of hundreds of thousands of individual molecules with nanometer precision in cells. In first proof-of-principle experiments, these techniques have been used to investigate the spatial arrangement of proteins in important structures such as the focal adhesion:
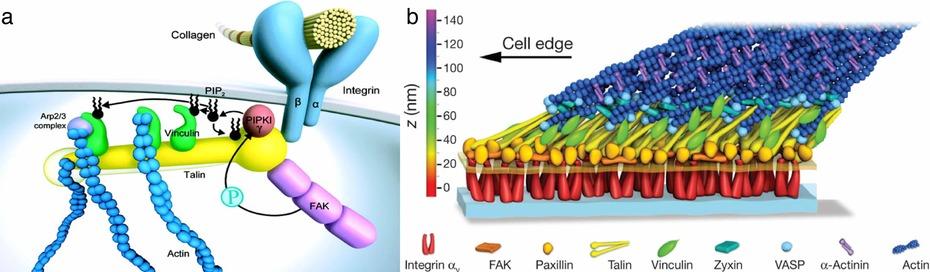
Figure 1: The focal adhesion. Shown are two data-based models for the focal adhesion. Most molecular players have been identified and many specific interactions have been characterized, often down to the atomic level. Based on this information a model for the focal adhesion is generated3 (a). What this modelcannot convey based on the present information is how the molecules are organized within the focal adhesion complex. The architecture of the focal adhesion complex was recently resolved using a novel single molecule localization based superresolution microscopy technique2 (b)
When Kanchanawong et al resolved the focal adhesion complex by single molecule superresolution microscopy, they could reveal striking organizational patterns within the thousands of molecules: i) they found that the entire complex is in itself oriented towards an extracellular cue, ii) the molecules in the complex are organized in a modular pattern where specific functions are concentrated in layers and iii) within the complex some molecules are always arranged in the same spatial orientation. These findings suggest that not only the composition of a multiprotein complex, but also the relative spatial orientation and organization of the components play an important role in generating the specific function.
We aim to develop single molecule superresolution microscopy to a state where it can readily be used as a readout of cell function. To that end, we develop novel labeling techniques and computational analysis routines. For example, we have pioneered the use of nanobodies in single molecule superresolution microscopy, allowing for higher resolution of cellular structures:
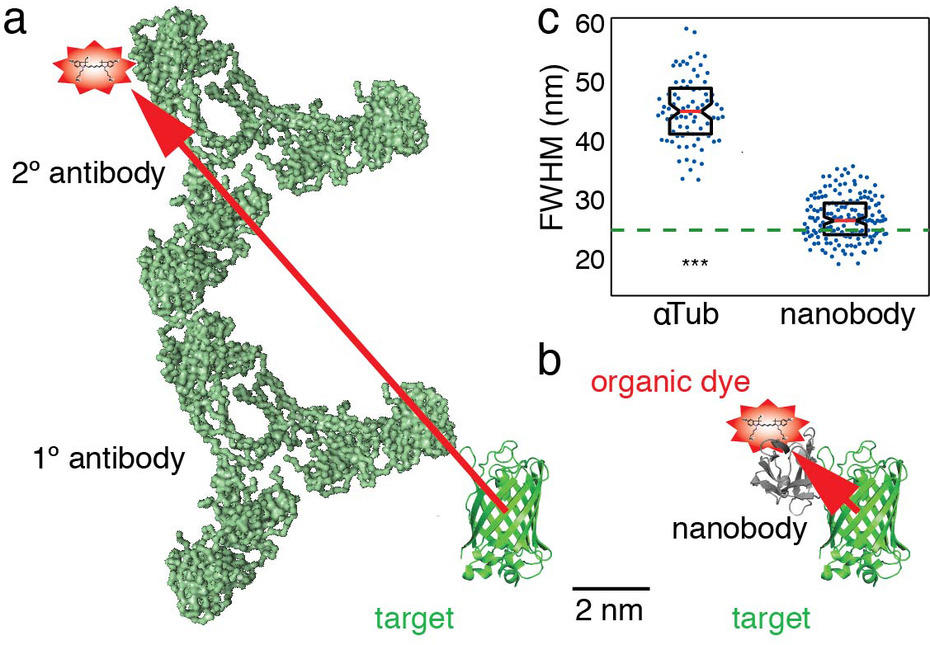
Figure 2: A simple and versatile novel labeling strategy. To achieve accuracies in localization below 10 nm in single molecule superresolution microscopy, organic dye labels are required. Traditionally, an organic dye is delivered to the target via a specific primary antibody and a dye-labeled secondary antibody (a). However, due to the size of the antibodies, the dye will be detected > 10 nm away from the target molecule (red arrow), adding a significant error in the detection of the target. We developed a novel approach using AF647 coupled nanobodies. Only 1.5 x 2.5 nm in size, these tiny binders can target any GFP-tagged structure in cells (b). This increases the resolution of the target. Anti-tubulin antibodies show a measured microtubule diameter of ~ 45 nm, compared to ~ 26 nm with nanobodies (c).
2. Understanding the organization and function of cellular septin filaments.
In progress.
3. Understanding how the plasma membrane is affected by membrane-cytoskeleton interactions.
In progress.