Johansson Group
Research interests
Light as an external signal has been shown to control many aspects of plant growth and development throughout the plant lifecycle including germination, de-etiolation, shade avoidance, chloroplast movements, phototropism, entrainment of the circadian clock and time of flowering. The seedling de-etiolation is one of the most dramatic response to any single external stimuli in nature and as such, it is not surprising that this response affects a large proportion of the plant transcriptome. This is largely mediated by the red and blue light receptors (phytochromes and cryptochromes). However, these photoreceptors do not regulate transcription directly but via a complex network of positive and negative factors of photomorphogenesis.
Over two decades ago, COP1 was identified as a major negative regulator of light signalling through its photomorphogenic phenotype in darkness. In the dark, COP1 acts as an E3 ubiquitin ligase targeting several positive factors of photomorphogenesis for degradation, most notably the bZIP transcription factor HY5. Upon light irradiation, COP1 activity is inhibited by the photoreceptors in addition to being excluded from the nucleus resulting in the accumulation of HY5 and consequently de-etiolation.
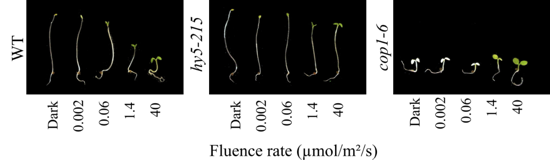
Photograph of seedlings grown for 7 days in darkness or increasing amounts of red light. Increasing light levels results in opening and expansion of the cotyledons, inhibition of hypocotyl elongation and chlorophyll accumulation. The hy5 mutant is hyposensitive to light while the cop1 mutant is photomorphogenic in darkness.
In the Johansson group we are interested in the molecular events taking place between the perception of light by the photoreceptors and the following transcriptional response during de-etiolation. We are working with a group of B-Box zinc finger proteins that physically interact with COP1 and/or HY5. As predicted by these interactions, these B-Box proteins do indeed modulate the light signalling pathway as manifested in the mutants by altered inhibition of hypocotyl elongation and accumulation of anthocyanin. Currently, we are examining the similarities and differences between these B-Box proteins and are working towards the mechanism by which they alter HY5 action.


